Team Christian Stolte LEAD UX DESIGN + VISUAL DESIGN Faygel Beren UX DESIGN Nathan Pearson BIOINFORMATICS LEAD Kevin Shi SOFTWARE ENGINEERING Gabrielle Bertier PROJECT MANAGEMENT Toby Bloom, John Greally PRINCIPAL INVESTIGATORS
Client NYCKidSeq Icahn School of Medicine at Mt. Sinai, Albert Einstein Medical College, New York Genome Center
“At least 70% of genetic tests are inconclusive — that is very frustrating.”
Physician from NYCKidSeq
The price of sequencing a genome has dropped to levels comparable with other medical tests. As a result, genetic testing has made its way into clinical practice. While genetic tests hold promise for many rare diseases, they also present problems:
• 75% of patients and their families are generally receptive to genetic testing, but they overestimate its practical usefulness.
• Most genetic test results fail to reveal a diagnosis.
• Physicians acknowledge a lack of knowledge about genetic testing and the available options.
The NYCKidSeq clinical trial in the Bronx would enroll 1130 children with neurologic, immunologic, or cardiac problems in genomic testing. Our main goal was to compare diagnostic rates between targeted gene panels and whole genome sequencing. A secondary goal was to develop a software tool that could improve the diagnostic rate by facilitating better communication between physicians and genetic testing labs.
The core idea for the functionality of the software came out of initial interviews with the genetic testing lab. When the lab scientists had more detailed information about the patients’ symptoms, they found related variants much more often.
In cases where the lab worked closely with a referring physician who was a geneticist, they were able to get feedback on candidate variants. They would also collect dozens of terms describing the patient’s condition. The lab could then feed those terms into their algorithms for analyzing the patient’s genome to identify variants that might be causal or helpful for diagnosis. In standard practice, the order form for a genetic test contained, at best, a handful of such terms. Contacting the physician was often impractical and fraught with delays. If you could get hold of them, chances were they didn’t understand genetics. We needed a way to help the lab communicate with a genetics nonexpert about specific genetic findings. It would also allow the clinicians to tell the lab details about their patients, without the added burden of writing long emails.
Design sprint to the rescue
We decided to hold a one-day stakeholder workshop that we organized around the idea of a design sprint. We invited a group representing caregivers, genetic testing laboratory, and research scientists: physicians, genetic counselors, laboratory geneticists, bioinformaticians, and biologists.
The agenda for the first session was to create user personas and map their journeys through the testing process.
The second session focused on pain points and goals. The fundamental needs to consider were:
Can we better engage the Clinician with the genetic testing results and exploration process?
Can we create an easy, worthwhile working experience or relationship between the clinician and the app?
Can we create an easy, worthwhile working experience or relationship between the clinician and the app?
In the next session we wanted to develop ideas for possible solutions, using the “Crazy 8s” method. Everyone had eight minutes to come up with eight rough ideas:

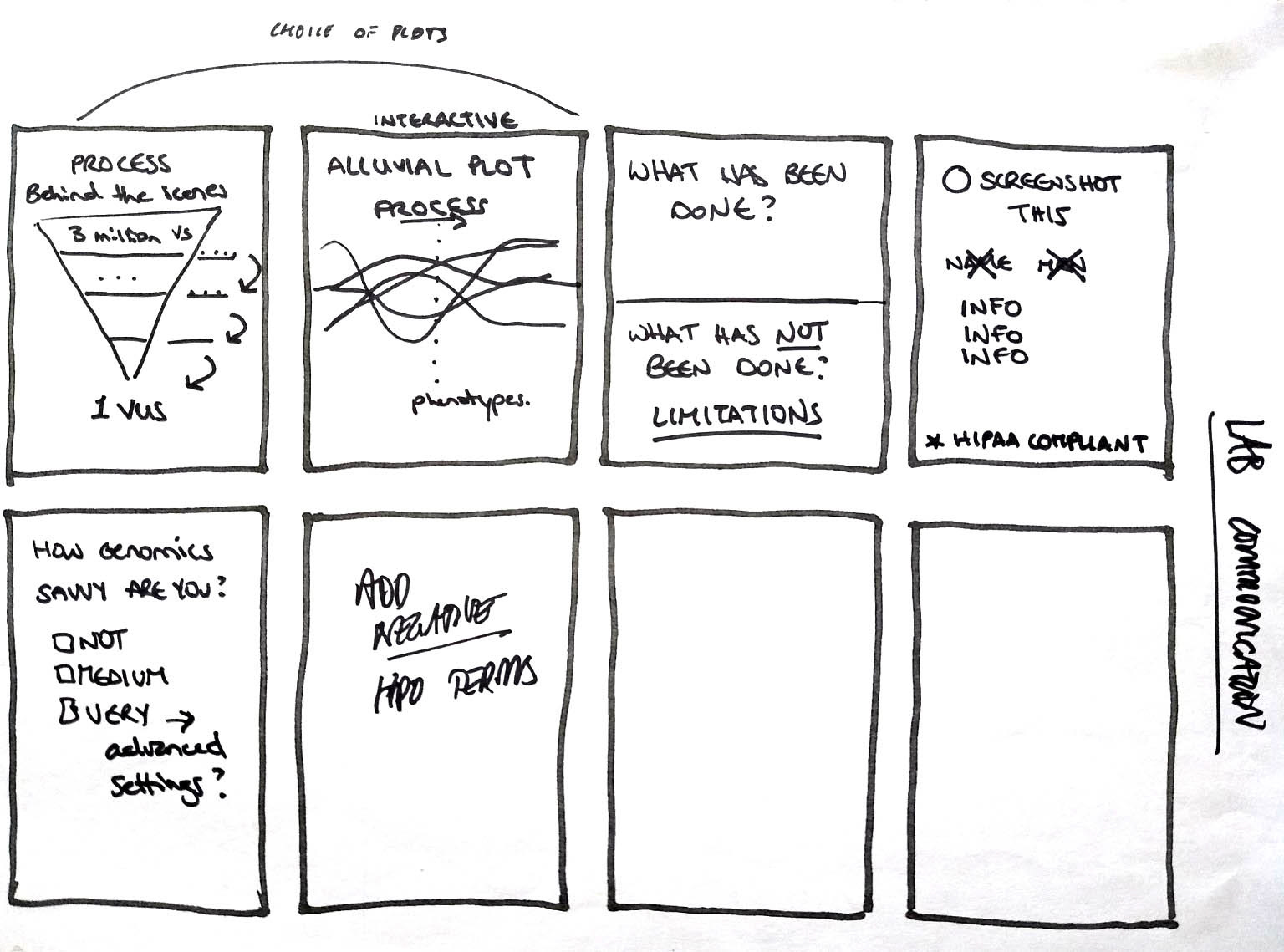



This brainstorming exercise produced a lot of interesting ideas; we hope to incorporate some of them into future versions of the tool. We identified three themes for further exploration:
• creating connections to online resources, based on results
• ranking results and potential causes
• exploring variability to expand the result space
• ranking results and potential causes
• exploring variability to expand the result space
We took those concepts to work on in the following session: prototype sketches.

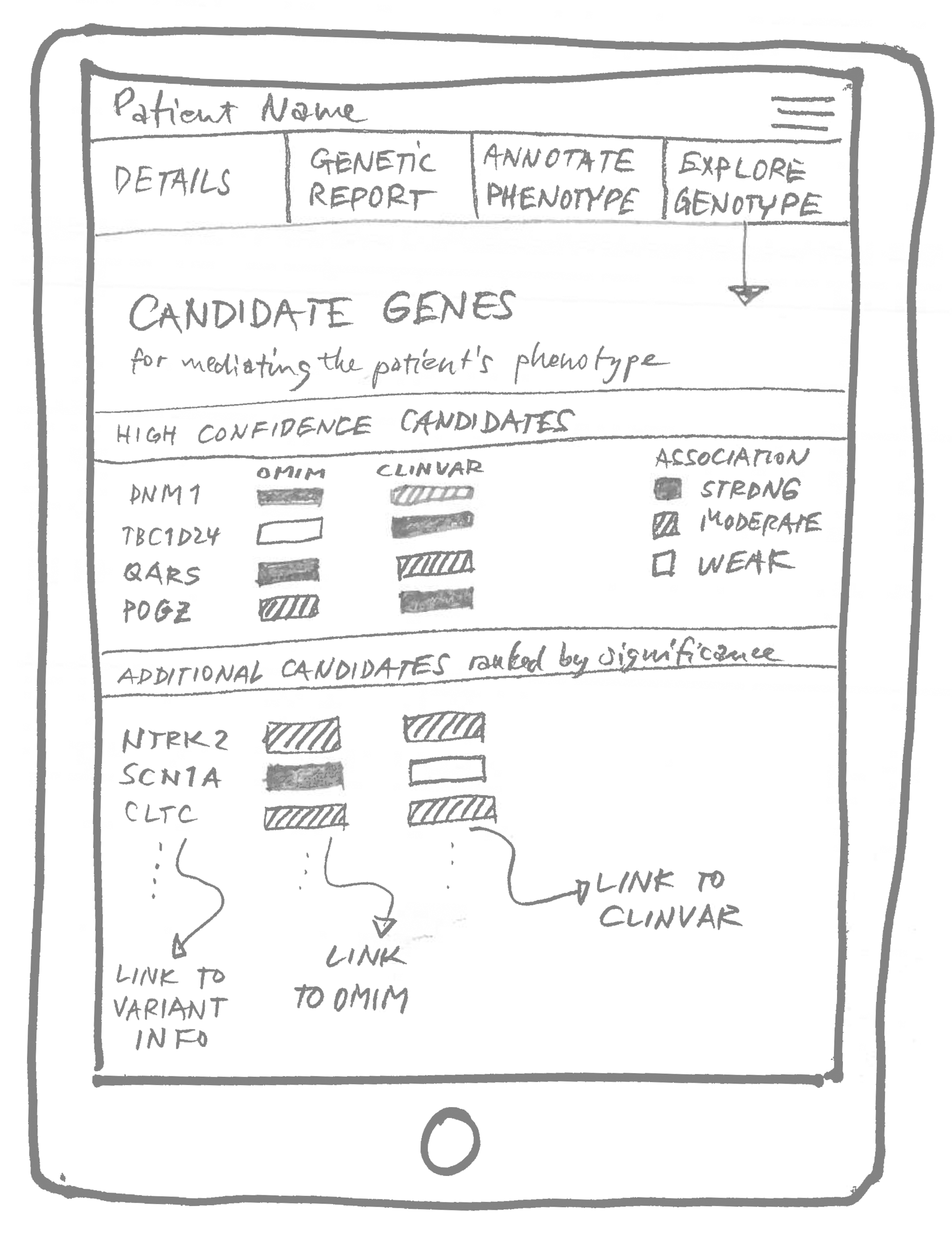
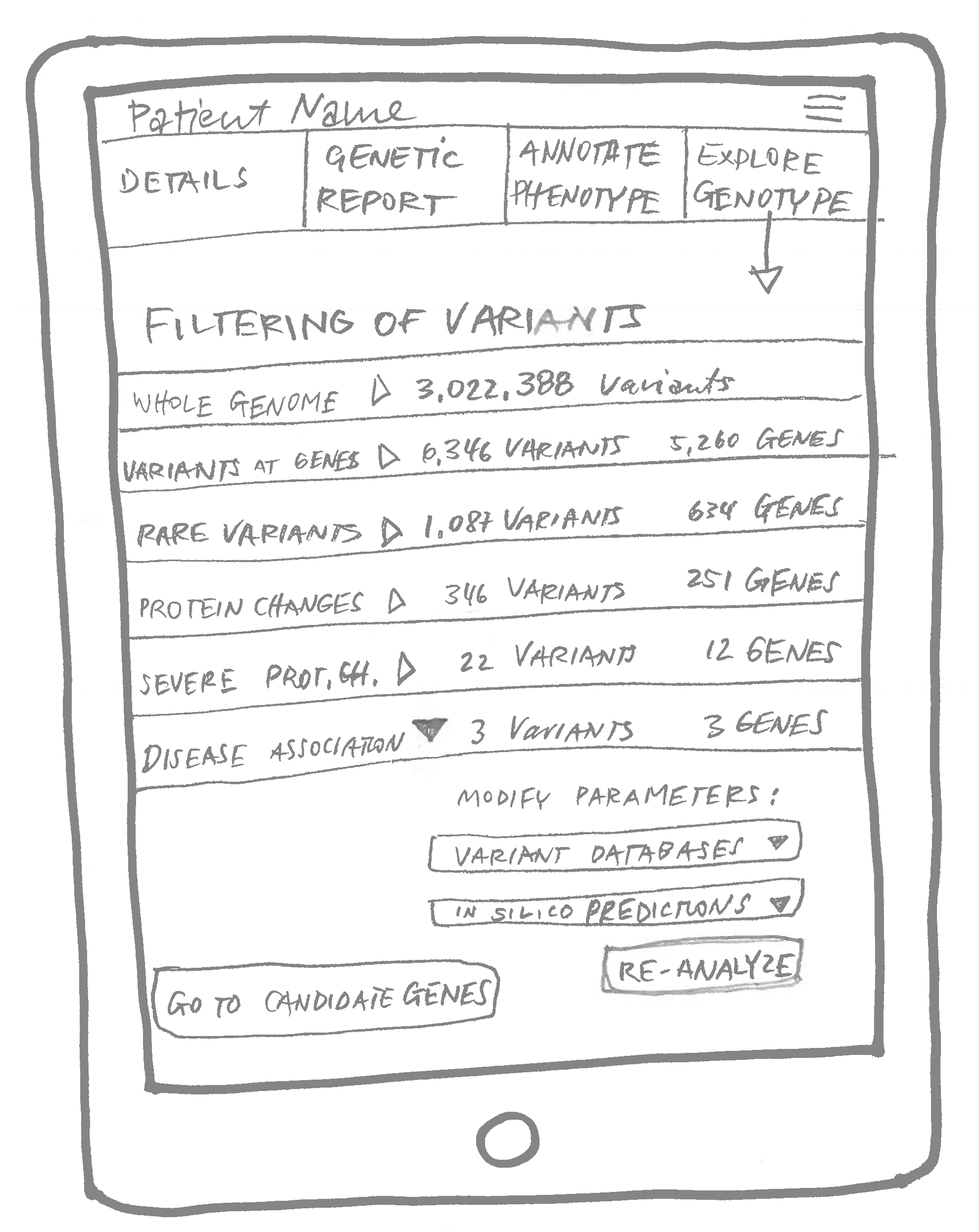
How to talk about genetics without mentioning genes
In our stakeholder interviews it had become clear: for many physicians genetics was a topic outside their comfort zone. So how could the lab get input from doctors about variants found in their patients’ genomes?
The answer lay dormant in online databases. If we could find the symptoms and traits associated with a genetic variant, we could ask questions that every doctor feels comfortable answering: “Does your patient have this symptom — yes, no, or maybe?”
As the first step of GenomeDiver’s workflow, the lab would upload the genomic data and terms describing the patient. The software would then find mutations in the data, called variants, that were potentially associated with the patient’s symptoms. Each variant, in turn, can point to one or more traits or symptoms. Those terms would then be presented to the clinician to decide if they were true for their patient.
The software would keep track of how the physician answered. It would calculate a score for each associated variant, then prepare a ranked list of variants for the lab. Along with it, the lab would receive the terms classified as “yes” and “maybe” to feed into their own software.
UI Designs
Our portal would primarily be accessed on laptops or desktop computers, occasionally on an iPad. We designed the UI with touch interactions in mind. Control elements would be sufficiently large, and links in the tables spaced far enough apart to avoid problems.
The following images illustrate the caregiver view:

A patient's record has been selected

Drag the terms into the right category; terms generated from the genetic data are marked with a double-helix icon.
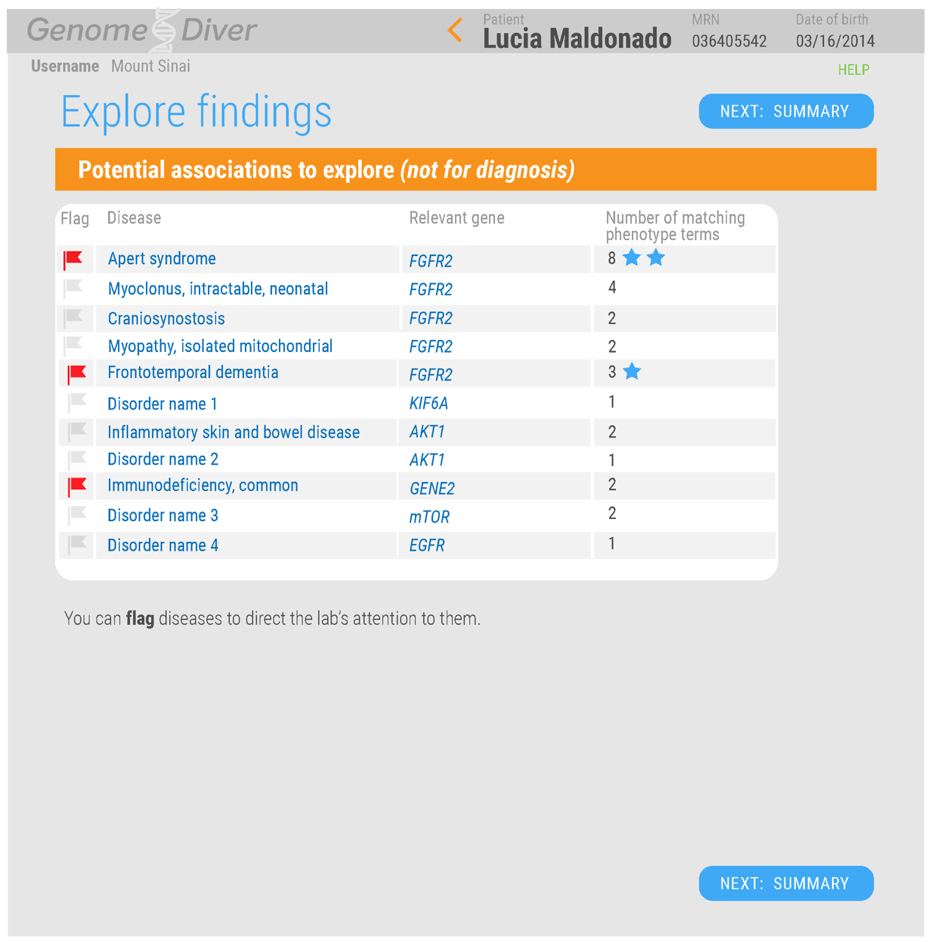
Flag diseases or click on the links to explore online resources
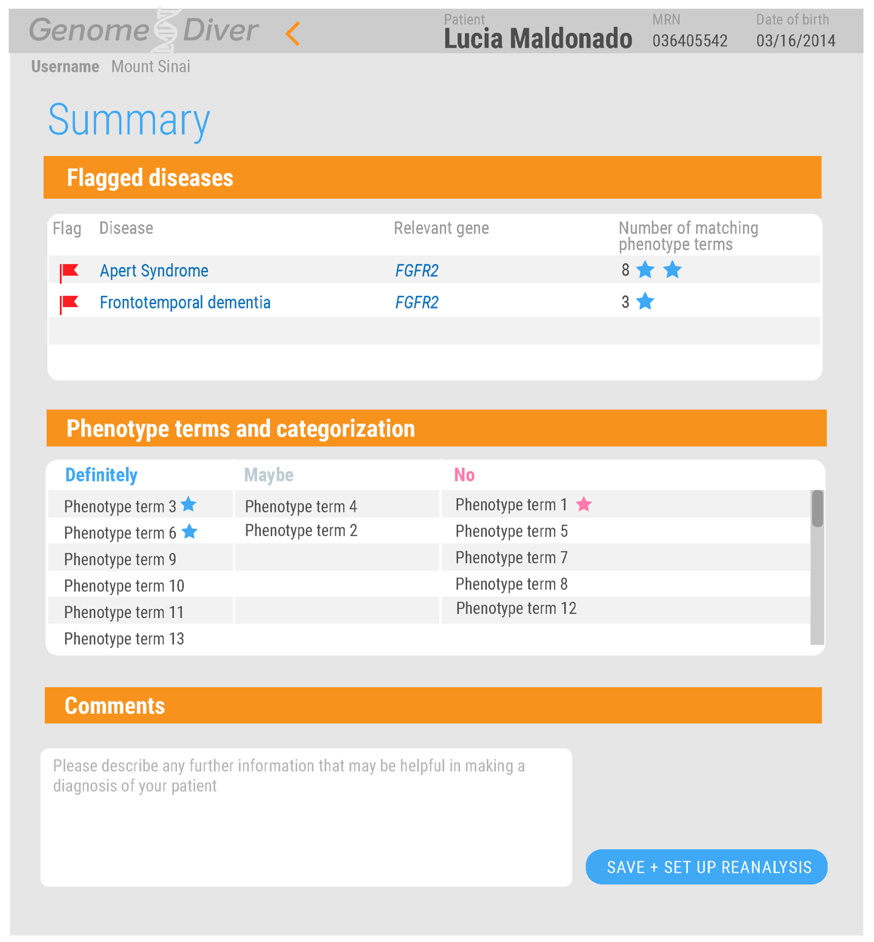
Summarize findings for the lab
We prepared a style guide to help the developers build the UI in a consistent manner.